MBH-Seg: Multi-class Brain Hemorrhage Segmentation in Non-contrast CT
News
[2024.7.24] We are extending the submission deadline for the 1st validation phase by one week.[2024.7.10] We add a new authentication method for participants registrations.
[2024.5.15] We opened the challenge website and released the training datasets.
Overview
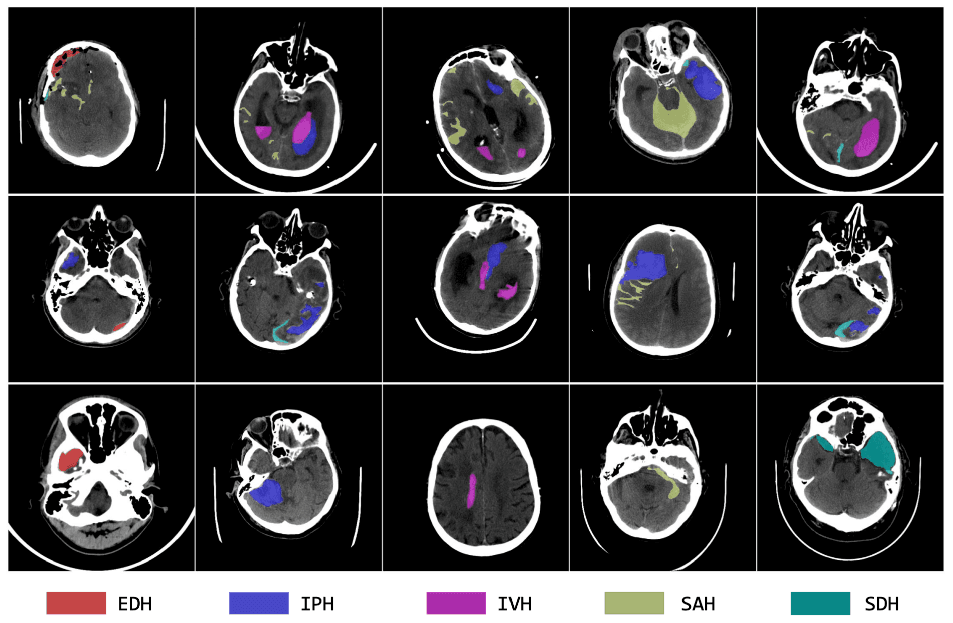
We invite you to participate in the first MBH-Seg Challenge. The focus of this year's challenge is to segment various types of brain hemorrhages base on non-contrast CT scans in a semi-supervised setting.Intracranial hemorrhage (ICH) encompasses bleeding within the skull or brain, manifesting in several types based on the anatomical location of the bleed relative to the brain and its surrounding membranes. These types include extradural hemorrhage (EDH), subdural hemorrhage (SDH), subarachnoid hemorrhage (SAH), intraparenchymal hemorrhage (IPH), and intraventricular hemorrhage (IVH). The causes of ICH are varied, ranging from trauma and vascular malformations to hypertension and venous thrombosis. Detection and characterization of ICH are typically performed using non-contrast CT scans, which provide crucial insights into the type and distribution of the hemorrhage. Accurate localization, quantification, and classification of these hemorrhages are vital for clinical management, as they help clinicians assess severity, predict patient outcomes, and determine appropriate treatment strategies. Despite advances in deep learning that have propelled the automation of medical image segmentation, the segmentation of different ICH classes remains a challenge, particularly due to the scarcity of public datasets with multi-class, pixel-level annotations. Most existing datasets focus on hemorrhage classification or single-class segmentation (foreground vs. background), thus limiting the development of detailed segmentation tools.
This challenge enhances a public ICH dataset to create a comprehensive 3D, multi-class ICH dataset. This dataset will feature precise pixel-level hemorrhage annotations alongside a significantly larger pool of unannotated data. Participants are encouraged to employ semi-supervised learning techniques to develop diagnostic tools that are not only more consistent and rapid but also more accurate. Such advancements could potentially improve patient outcomes by reducing mortality and furthering the application of deep learning in medical imaging.
Prizes
1. Cash Awards:First Prize: $800
Second Prize: $400
Third Prize: $200
2. Certificates for the top-rank teams.
3. The top-10 teams will be invited to give oral presentations at the MICCAI 2024 Event (either virtually or in person).
4. Co-author of the challenge paper, which will be submitted to a top journal (MedIA/TMI).
Schedule
| Event | Timeline | ||||
|---|---|---|---|---|---|
| Release of training data | May 15th, 2024 | ||||
| Submission for the 1st-Validation phase | From July 29th, 2024 to August 22nd, 2024 | ||||
| Submission for the 2nd-Testing phase | From August 22nd, 2024 to September 8th, 2024 | ||||
| Announcement of invited presentations | September 28th, 2024 | ||||
| Announcement of final leaderboards | October 6th, 2024 | ||||
Dataset
Citations
1. Wu, Biao, Yutong Xie, Zeyu Zhang, Jinchao Ge, Kaspar Yaxley, Suzan Bahadir, Qi Wu, Yifan Liu, and Minh-Son To. "BHSD: A 3D Multi-class Brain Hemorrhage Segmentation Dataset." In International Workshop on Machine Learning in Medical Imaging, pp. 147-156. Cham: Springer Nature Switzerland, 2023.
2. Anouk Stein, MD, Carol Wu, Chris Carr, George Shih, Jayashree Kalpathy-Cramer, Julia Elliott, kalpathy, Luciano Prevedello, Marc Kohli, MD, Matt Lungren, Phil Culliton, Robyn Ball, Safwan Halabi MD. (2019). RSNA Intracranial Hemorrhage Detection. Kaggle. https://kaggle.com/competitions/rsna-intracranial-hemorrhage-detection
2. Anouk Stein, MD, Carol Wu, Chris Carr, George Shih, Jayashree Kalpathy-Cramer, Julia Elliott, kalpathy, Luciano Prevedello, Marc Kohli, MD, Matt Lungren, Phil Culliton, Robyn Ball, Safwan Halabi MD. (2019). RSNA Intracranial Hemorrhage Detection. Kaggle. https://kaggle.com/competitions/rsna-intracranial-hemorrhage-detection
Evaluation
Metric(s)
1. Dice Similarity Coefficient (Dice): Dice measures the overlap between the predicted segmentation and the ground truth at the pixel-level. It's highly relevant for medical image segmentation to ensure accurate delineation of affected areas.
2. Sensitivity: This metric assesses the class-level true positive rate, which refers to the test's ability to correctly detect ill patients out of those who do have the condition.
3. Specificity: This metric assesses the class-level true negative rate, which refers to the test's ability to correctly reject healthy patients without a condition.
For the final ranking, metrics will be aggregated for the Challenge Score (CS) as:
CS = 0.5 * Dice + 0.25 * Sensitivity + 0.25 * Specificity
Submission
Coming soon...Terms and Conditions
1. Only automatic methods are allowed
2. No additional data is allowed. We will require all participants to provide extensive documentation on their development processes, including data and methods used. Top-ranked participants must submit their training code and checkpoints for verification. Publicly available pre-trained models are allowed, ensuring a level playing field and transparency in the competition's evaluation process.
3. The top-10 teams will be notified two weeks before the challenge day to prepare their presentations. Final results and awards will be announced on the challenge day.
4. Independent Publications: Participating teams are allowed to publish their results separately. However, this should be done in a manner that respects the collective efforts of the challenge.
5. Embargo Period: An embargo period allows the challenge organizers to publish a comprehensive challenge paper first. During this period, individual teams are encouraged to refrain from publishing their complete findings independently.
2. No additional data is allowed. We will require all participants to provide extensive documentation on their development processes, including data and methods used. Top-ranked participants must submit their training code and checkpoints for verification. Publicly available pre-trained models are allowed, ensuring a level playing field and transparency in the competition's evaluation process.
3. The top-10 teams will be notified two weeks before the challenge day to prepare their presentations. Final results and awards will be announced on the challenge day.
4. Independent Publications: Participating teams are allowed to publish their results separately. However, this should be done in a manner that respects the collective efforts of the challenge.
5. Embargo Period: An embargo period allows the challenge organizers to publish a comprehensive challenge paper first. During this period, individual teams are encouraged to refrain from publishing their complete findings independently.
Community
Organizers
•Yutong Xie, Researcher, Australian Institute for Machine Learning (AIML), University of Adelaide, Australia
•Minh-Son To, Medical Practitioner, Flinders Health and Medical Research Institute, Flinders University, Australia
•Chenyu Wang, Senior Lecturer, Brain and Mind Centre, University of Sydney, Australia
•Dongang Wang, Researcher, University of Sydney, Australia
•Zhibin Liao, Researcher, Australian Institute for Machine Learning (AIML), University of Adelaide, Australia
Advisor
•Qi Wu, Assoiate Professor, Australian Institute for Machine Learning (AIML), University of Adelaide, Australia
•Johan Verjans, Clinician-scientist, Royal Adelaide Hospital; Associate Professor, Australian Institute for Machine Learning (AIML), University of Adelaide, Australia
•Yong Xia, Professor, Northwestern Polytechnical University, China
Contributors
•Biao Wu, Student, University of Adelaide, Australia•Siqi Chen, Research Assistant, Australian Institute for Machine Learning (AIML), University of Adelaide, Australia
•Vu Minh Hieu Phan, Researcher, Australian Institute for Machine Learning (AIML), University of Adelaide, Australia
•Yifan Liu, Lecturer, Australian Institute for Machine Learning (AIML), University of Adelaide, Australia
Organized By





Contact Us
Please contact us if you have any enquires through our Google Form